OpenAge.
An open-weight
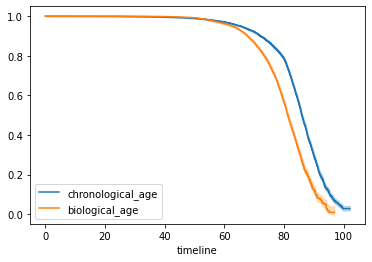
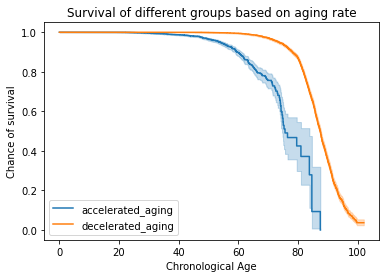
biological aging clock
validated against
actual death.
on most of biology.
You can on aging."
Nikhil Yadala · Healome One Inc.
github.com/Healome/OpenAge · OpenAgeAI.com

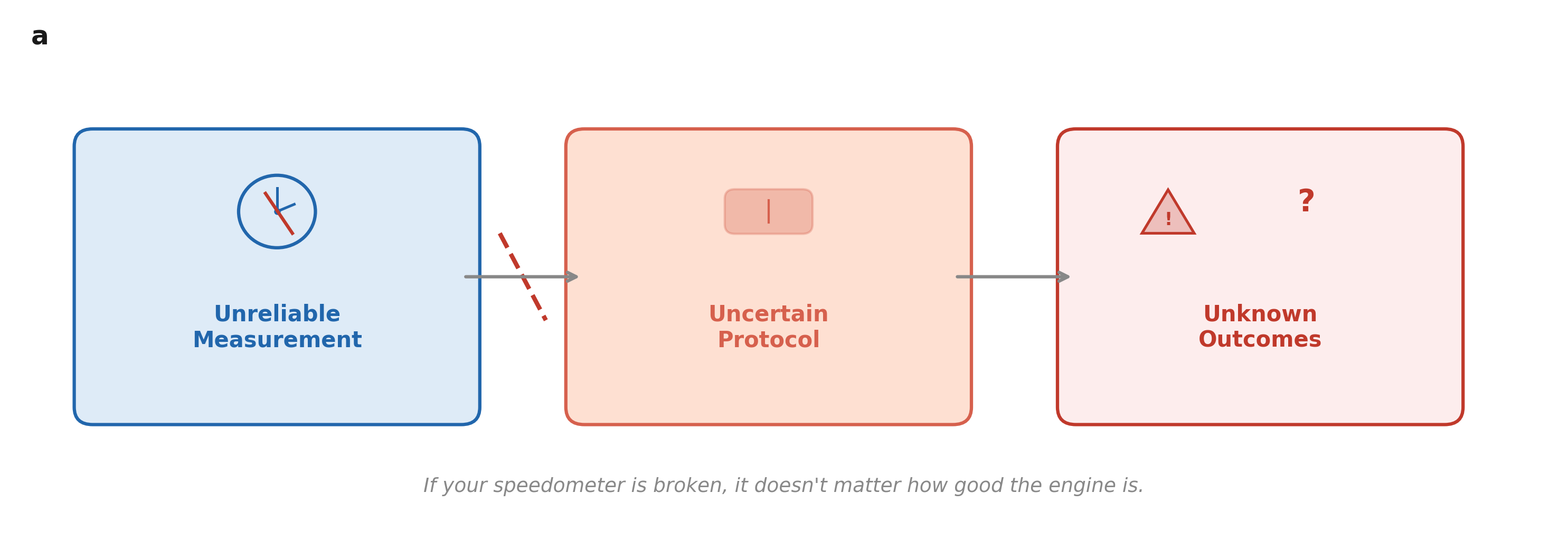
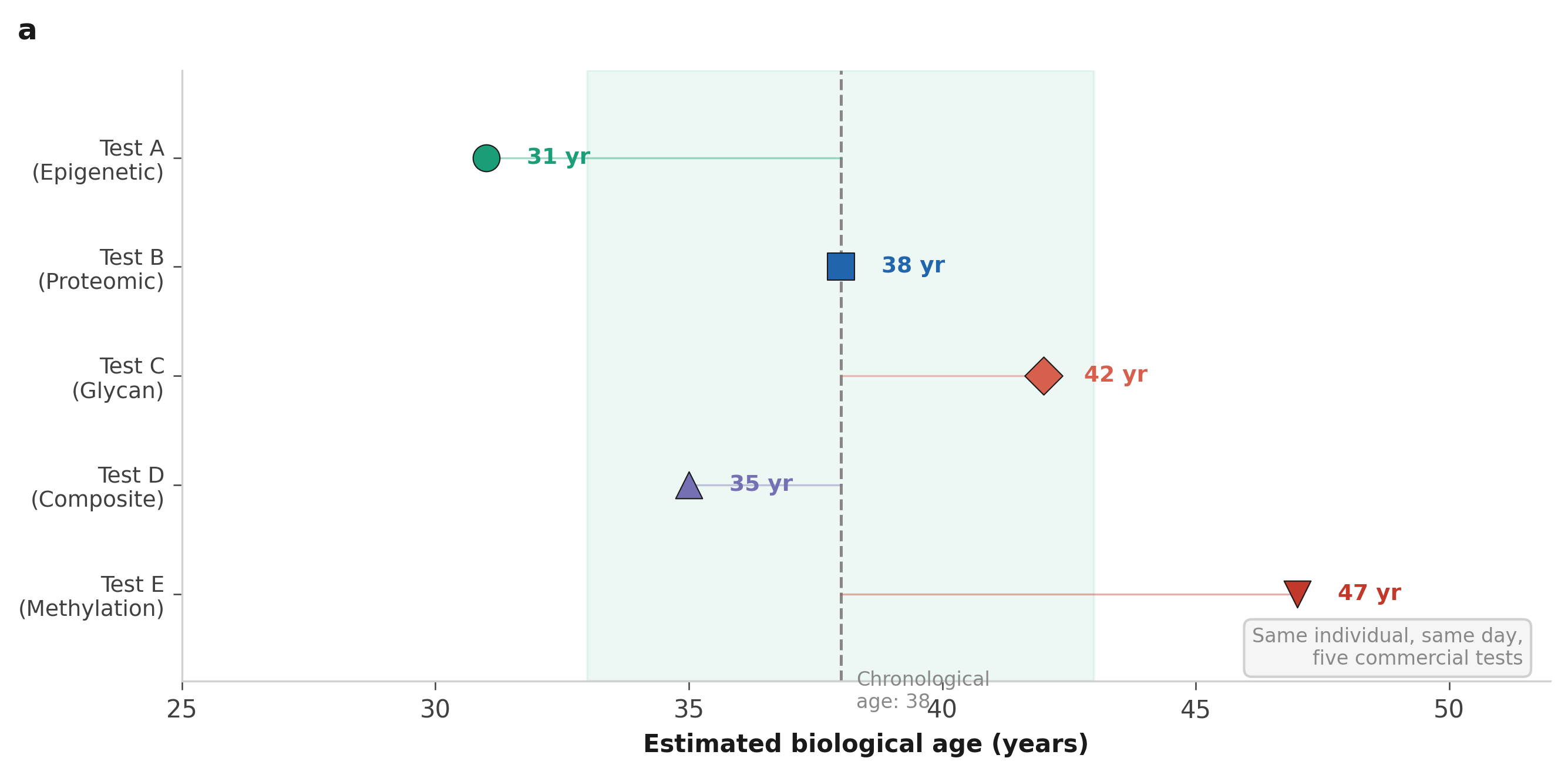
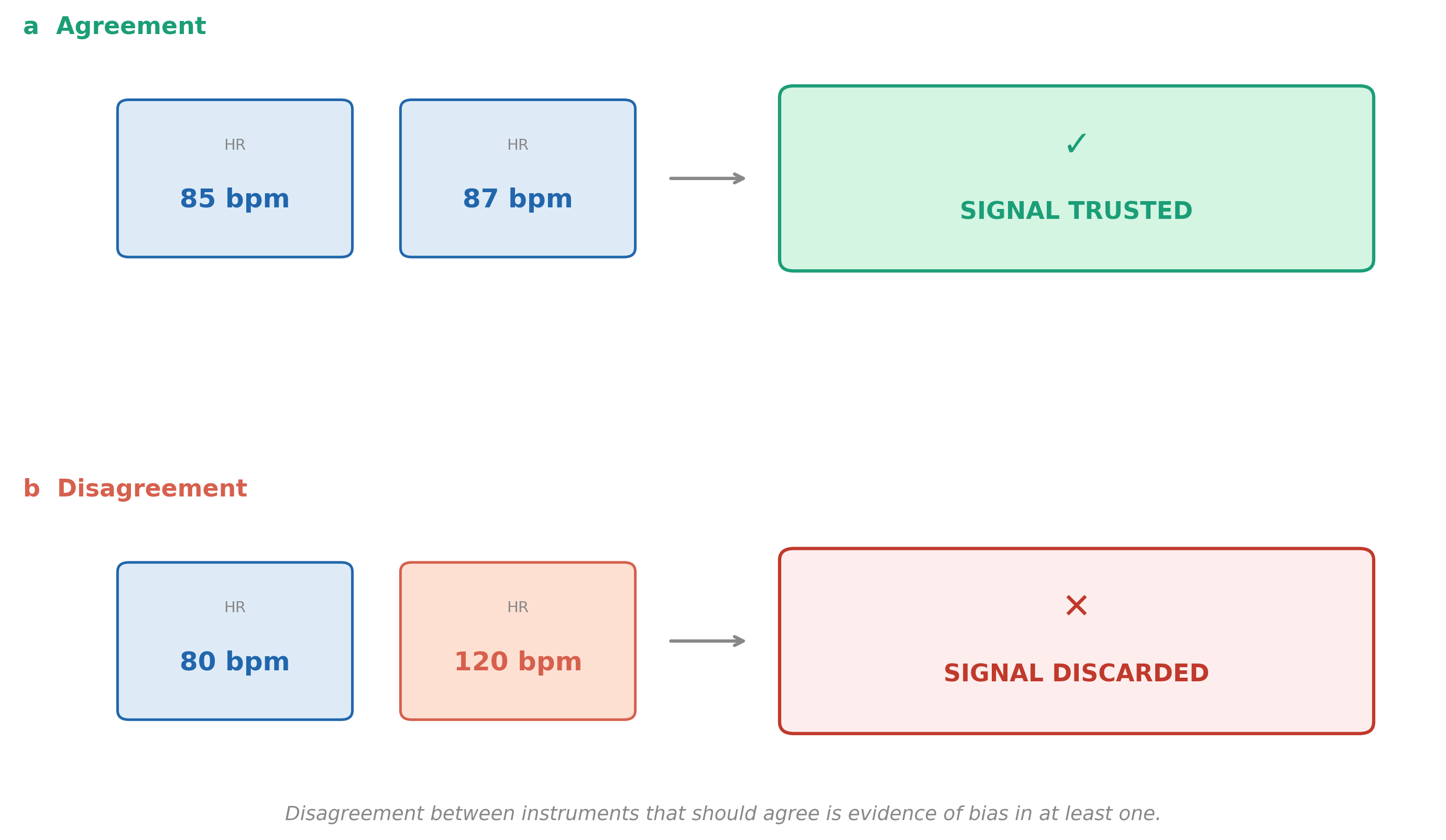
Different tests. Different answers.